Fluorescence microscopy
Fluorescence microscopy is a well-suited technique for examining fixed and living specimens and both topological features as well as fluorescence intensity variations can be evaluated in a quantitative manner. Fluorescent compounds emit light in a defined spectral range when excited with light of a specific wavelength. Fluorescence imaging makes use of this phenomenon and allows for the selective and specific detection of molecules at small concentrations with a good signal-to-noise ratio. Furthermore, detection of the signal by confocal microscopy allows to acquire images with good resolution in the focal plane, suppressing any signal arising from out-of-focus regions. In our lab, we use different fluorescence microscopy imaging techniques, e.g. wide-field, confocal laser scanning microscopy and automated 3D screening. In combination with image analysis, we detect and quantify the properties of different molecules such as DNA, RNA or proteins at specific cellular locations, which aids to deduce their functions and underlying molecular mechanisms.
|
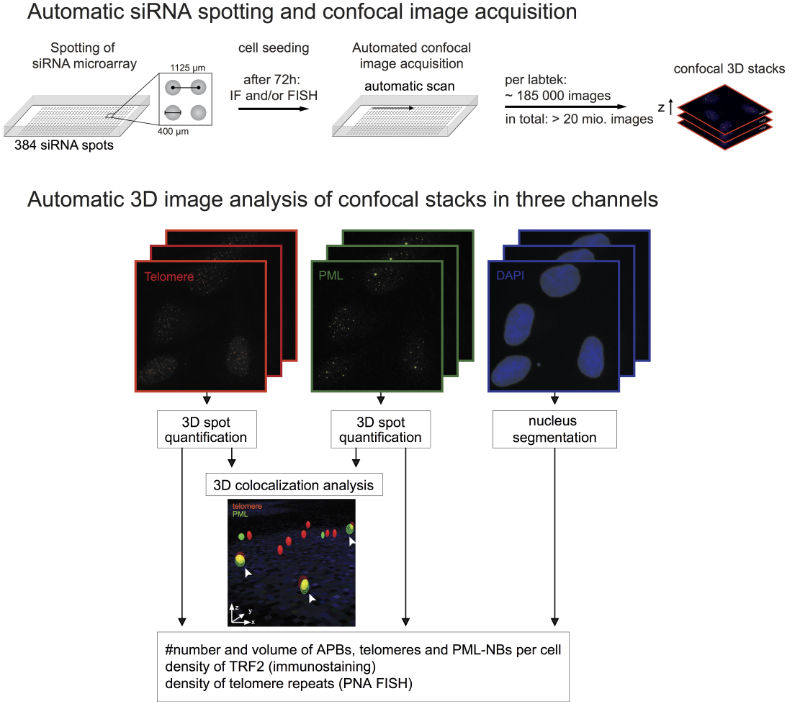
| Automated confocal microscopy for screening. The workflow for implementation of an siRNA based screening approach together with the subsequent automatic image analysis is shown. |
Apart from imaging, fluorescence microscopy-based techniques are essential tools for measuring the dynamics of molecules in living cells to approach a variety of biological questions. The main fluorescence fluctuation microscopy techniques applied in our lab are fluorescence correlation spectroscopy (FCS) and fluorescence bleaching techniques (e.g. FRAP). A combination of these complementary methods allows for a comprehensive analysis on multiple length and time scales.
Key references
Rademacher A, Erdel F, Trojanowski J, Rippe K (2020) Tracing reversible light-induced chromatin binding with near-infrared fluorescent proteins. Meth Mol Biol 2173, 171-188. doi: 10.1007/978-1-0716-0755-8_12 | Abstract | Reprint (0.8 MB) | Article metrics
Trojanowski J, Rademacher A, Erdel F & Rippe K (2019) Light-induced transcription activation for time-lapse microscopy experiments in living cells. Methods Mol Biol 2028, 251-270. doi: 10.1007/978-1-4939-9674-2_17 | Abstract | Reprint (0.8 MB)
Gunkel M, Chung I, Wörz S, Deeg KI, Simon R, Sauter G, Jones DT, Korshunov A, Rohr K, Erfle H & Rippe K (2017). Quantification of telomere features in tumor tissue sections by an automated 3D imaging-based workflow. Methods 114, 60-73. doi: 10.1016/j.ymeth.2016.09.014 | Abstract | Reprint (7.3 MB) | Article metrics.